Regulation of OsMYB57 gene expression in rice and its allelopathic suppression to weed
-
摘要: 转录因子MYB57能够调控水稻化感抑草能力。为进一步研究调控OsMYB57基因表达的上游因子, 以化感水稻‘PI312777’和非化感水稻‘Lemont’为对象, 分离鉴定了OsMYB57基因启动子的转录因子。结果显示, OsMYB57基因表达受转录因子basic helix-loop-helix protein 009 (bHLH009, 基因: OsMYC2)、LOC_Os04g32590.1 (基因: Os04g0397500)、LOC_Os02g31160.1 (基因: Os02g0516800)以及转录调节因子 LOC_Os03g25430.1 (基因: Os03g0370250)、LOC_Os03g50110.1 (基因: Os03g0708800)等的调控。稗草胁迫下的‘PI312777’和‘Lemont’中的这些基因上调表达。bHLH009 (OsMYC2)是茉莉酸信号途径的调节因子, 外源添加0.05 mmol∙L−1茉莉酸甲酯(MeJA)不同程度提高了‘PI312777’和‘Lemont’中的OsMYC2、Os04g0397500、Os02g0516800、Os03g0370250和
Os03g0708800基因的表达, 水稻根系分泌液对稗草的化感抑制率提高; 化感水稻‘PI312777’根系分泌液对稗草地上和地下部生物量的抑制率分别提高25.82%和17.01%; 而非化感水稻‘Lemont’根系分泌液的抑制率则分别提高5.30%和15.36%。本研究表明, OsMYB57基因表达受bHLH009 (MYC2)等的转录调控, 适宜浓度的茉莉酸甲酯则能够促进OsMYC2的表达, 进而调控提高OsMYB57表达, 增强水稻化感抑草能力。 -
关键词:
- 化感作用 /
- 茉莉酸甲酯 /
- 转录调控 /
- 水稻 /
- 转录因子bHLH009
Abstract: Rice allelopathy is a natural inhibitory phenomenon of some rice varieties against surrounding weeds. It is an eco-friendly method to control weeds, and it has become a popular research topic in the development of sustainable agriculture. Allelopathy is a quantitative trait regulated by a number of genes that are responsive to environmental changes. Previous studies have indicated that increasing the gene expression of the R2R3-MYB transcription factor MYB57 increases the expression of genes involved in the synthesis of phenolic acid allelochemicals, thereby enhancing the allelopathic effect of rice. This demonstrated that the transcription factor MYB57 efficiently regulates allelopathic weed suppression in rice; however, the regulatory network for MYB57 remains unknown. To further explore the transcriptional factors that regulate the gene expression of OsMYB57, an allelopathic rice accession (‘PI312777’) and a non-allelopathic accession (‘Lemont’) were used. The amplification of the promoter region of the OsMYB57 gene in ‘PI312777’ and ‘Lemont’ was conducted using specific biotin-labeled primers at the 5′ end. The proteins binding to the OsMYB57 gene promoter were co-precipitated using the DNA pull-down technique. Identification of these proteins using mass spectrometry showed that the gene expression of OsMYB57 was regulated by transcription factors (basic helix-loop-helix protein 009 [bHLH009], LOC_Os04g32590.1, and LOC_Os02g31160.1) and transcription regulators (LOC_Os03g25430.1 and LOC_Os03g50110.1), which were encoded by the OsMYC2, Os04g0397500, Os02g0516800, Os03g0370250, and Os03g0708800 genes, respectively. The transcript level of these genes was up-regulated in ‘PI312777’ and ‘Lemont’ when they were co-cultured with barnyardgrass (Echinochloa crusgalli L.) for 1, 3, 5, and 7 days; more specifically, the expression of OsMYC2, Os02g0516800, and Os03g0708800 was significantly up-regulated in rice that was co-cultured with barnyardgrass for 5 days when compared to control groups. Among these proteins, basic helix-loop-helix protein 009 is a positive regulator of the jasmonate (JA) signaling pathway. The gene expression level of OsMYC2, Os04g0397500, Os02g0516800, Os03g0370250, and Os03g0708800 in ‘PI312777’ and ‘Lemont’ was up-regulated after the rice was treated with 0.05 mmol∙L−1 of exogenous methyl jasmonate (MeJA), which resulted in an increased allelopathic suppression to barnyardgrass. Results of western blotting demonstrated that the protein expression levels of MYB57, MAPK11, and PAL2;3 in ‘PI312777’ and ‘Lemont’ were increased under the induction of MeJA. Furthermore, when compared with the control group of barnyardgrass treated independently with MeJA, the root exudates of ‘PI312777’ and ‘Lemont’ induced by 0.05, 0.10, and 0.20 mmol∙L−1 of MeJA exhibited an enhanced growth inhibitory ratio, and the inhibitory effect of the root exudates of ‘PI312777’ induced by MeJA was higher than that of ‘Lemont’. The inhibitory ratios of ‘PI312777’ exudates induced by 0.05 mmol∙L−1 of MeJA on the plant height and root length of barnyardgrass increased by 25.82% and 17.01%, respectively; while the same treatment using ‘Lemont’ root exudates showed a 5.30% and 15.36% increased inhibitory ratio on barnyardgrass. These results indicated that the gene expression of OsMYB57 was transcriptionally regulated by bHLH009 (MYC2), and that a suitable concentration of MeJA promoted the gene expression of OsMYC2, thereby enhancing OsMYB57 expression and resulting in an increase in MYB57, MAPK11, and PAL2;3 expression, which led to systematically up-regulated and improved rice allelopathic weed suppression.-
Key words:
- Allelopathy /
- Methyl jasmonate /
- Transcriptional regulation /
- Rice /
- Transcription factor bHLH009
-
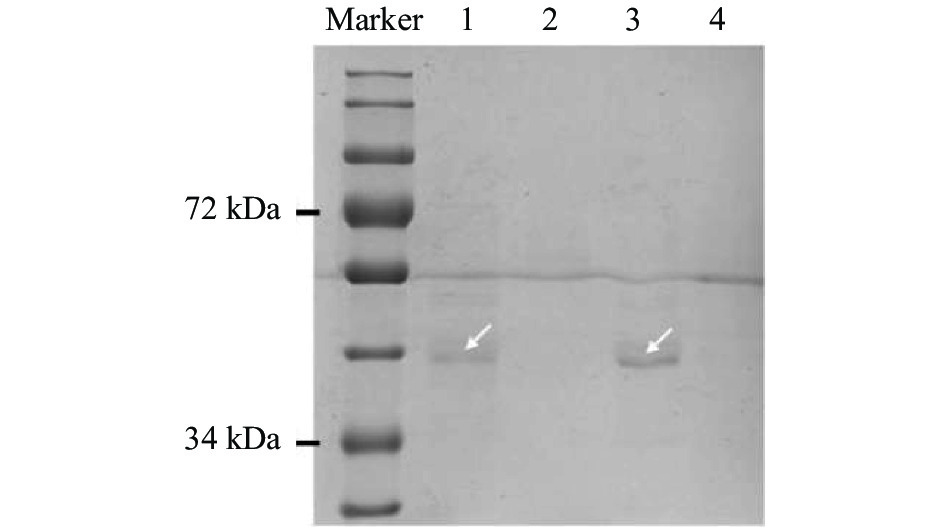
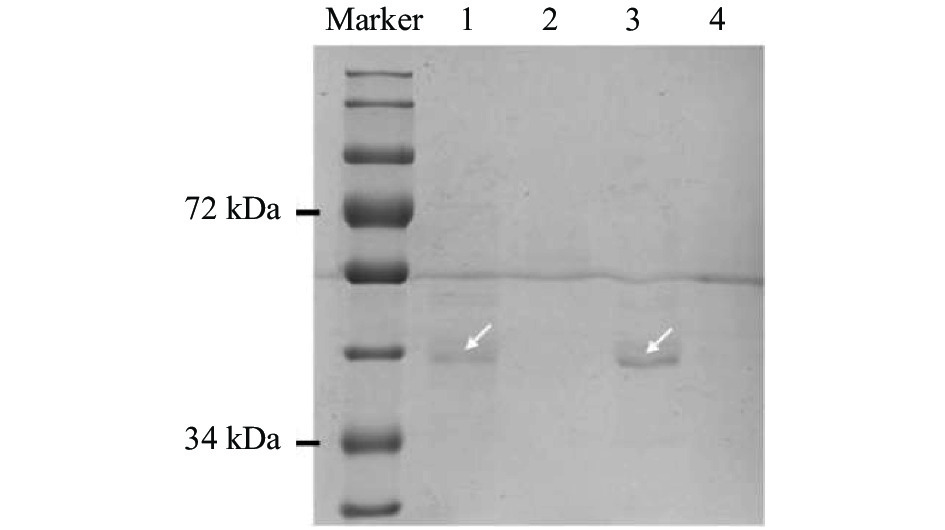
图 1 ‘PI312777’和‘Lemont’水稻中OsMYB57基因启动子的结合蛋白
Marker: 标准蛋白Marker; 泳道1: 稗草胁迫下‘PI312777’的结合蛋白; 泳道2: ‘PI312777’ 的阴性对照; 泳道3: 稗草胁迫下‘Lemont’的结合蛋白; 泳道4: ‘Lemont’的阴性对照; 箭头所指为结合蛋白。Marker: protein ladder; Lane 1: binding proteins from ‘PI312777’ co-cultured with barnyardgrass; Lane 2: negative control of ‘PI312777’; Lane 3: binding proteins from ‘Lemont’ co-cultured with barnyardgrass; Lane 4: negative control of ‘Lemont’. Arrow indicates the binding proteins.
Figure 1. Proteins binding on the promoter of OsMYB57 from rice varieties ‘PI312777’ and ‘Lemont’
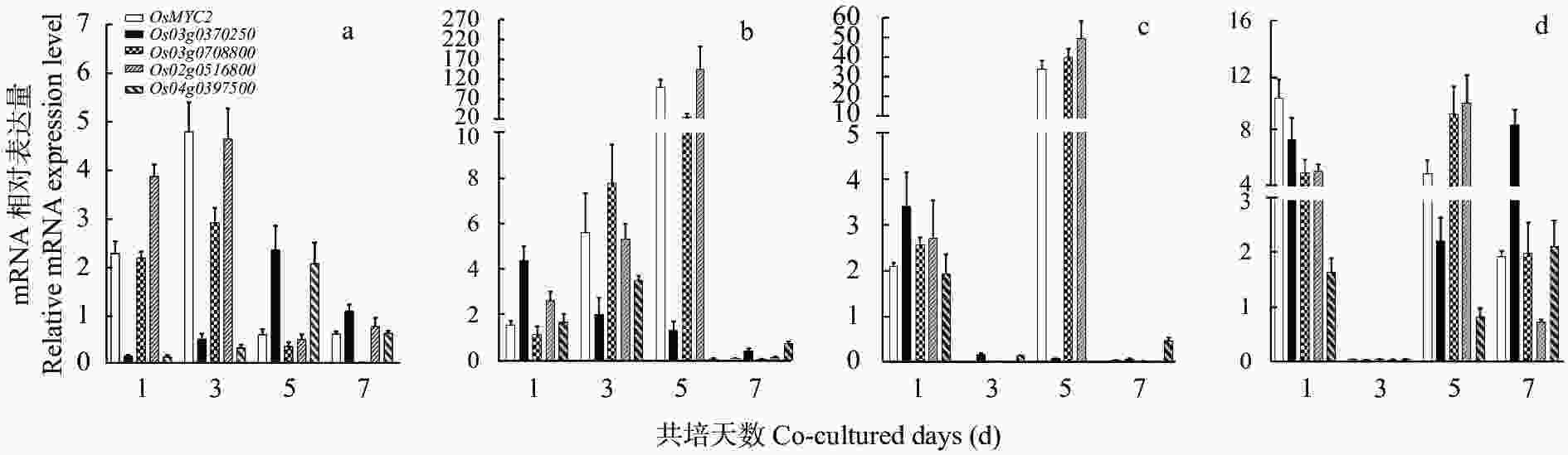
图 2 稗草胁迫下‘PI312777’和‘Lemont’水稻中OsMYB57基因转录调控因子的动态表达
a: ‘PI312777’叶中基因的表达; b: ‘PI312777’根中基因的表达; c: ‘Lemont’叶中基因的表达; d: ‘Lemont’根中基因的表达。a: gene expression in ‘PI312777’ leaves; b: gene expression in ‘PI312777’ roots; c: gene expression in ‘Lemont’ leaves; d: gene expression in ‘Lemont’ roots.
Figure 2. Dynamic gene expression levels of transcriptional regulators of OsMYB57 in rice varieties ‘PI312777’ and ‘Lemont’ co-cultured with barnyardgrass
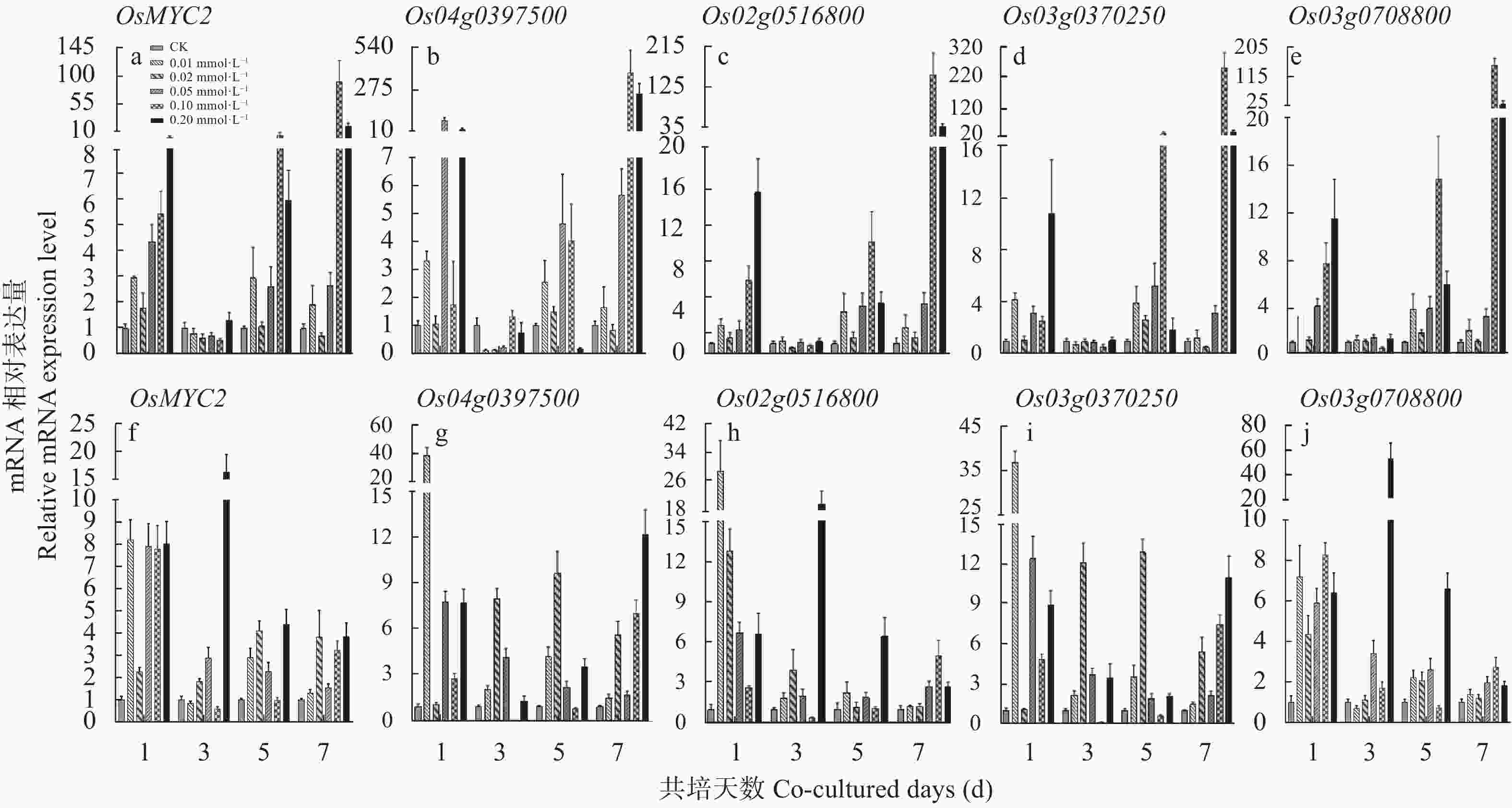
图 3 不同浓度茉莉酸甲酯处理不同时间‘PI312777’和‘Lemont’水稻根系中基因的动态表达
a−e: ‘PI312777’水稻根系中5个基因的表达; f−j: ‘Lemont’水稻根系中5个基因的表达。a−e: gene expression level in the root of ‘PI312777’; f−j: dynamic gene expression level in the root of ‘Lemont’.
Figure 3. Dynamics of gene expression level in the root of ‘PI312777’ and ‘Lemont’ under treatments of different concentrations of methyl jasmonate
图 4 不同浓度茉莉酸甲酯处理不同时间‘PI312777’和‘Lemont’水稻叶片中基因的动态表达
a−e: ‘PI312777’水稻叶片中5个基因的表达; f−j: ‘Lemont’水稻叶片中5个基因的表达。a−e: gene expression level in the leaves of ‘PI312777’; f−j: gene expression level in the leaves of ‘Lemont’.
Figure 4. Dynamics of gene expression level in the leaves of ‘PI312777’ and ‘Lemont’ under treatments of different concentrations of methyl jasmonate
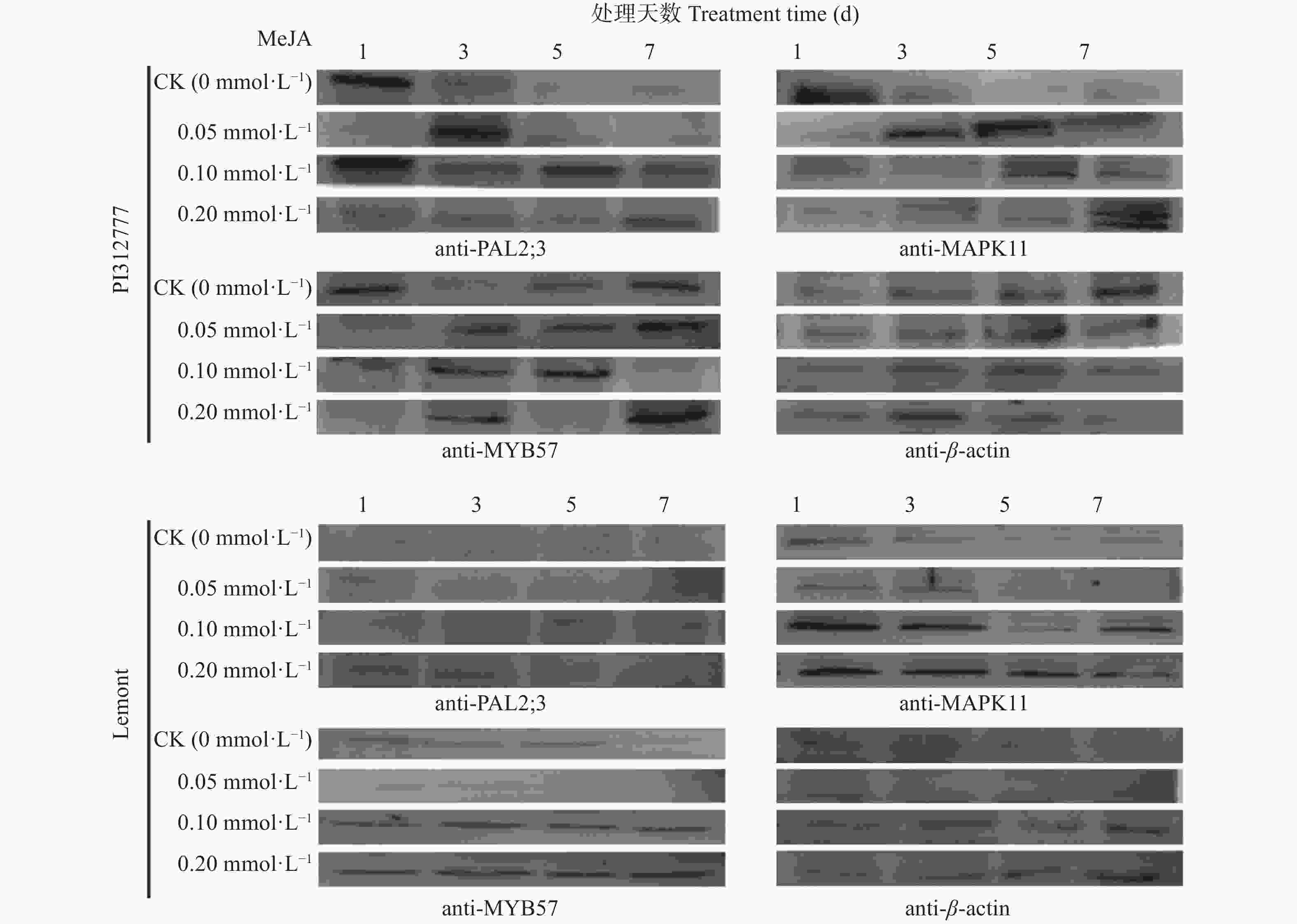
图 5 不同浓度茉莉酸处理不同时间‘PI312777’和‘Lemont’水稻中MYB57、PAL2;3和MAPK11蛋白的表达量
anti-PAL2;3: PAL2;3抗体; anti-MAPK11: MAPK11抗体; anti-MYB57: MYB57抗体; anti-β-actin: β-肌动蛋白抗体。anti-PAL2;3: antibody of PAL2;3; anti-MAPK11: antibody of MAPK11; anti-MYB57: antibody of MYB57; anti-β-actin: antibody of β-actin.
Figure 5. Protein expression of MYB57, PAL2;3 and MAPK11 in ‘PI312777’ and ‘Lemont’ under treatments of different concentrations of methyl jasmonate for different days
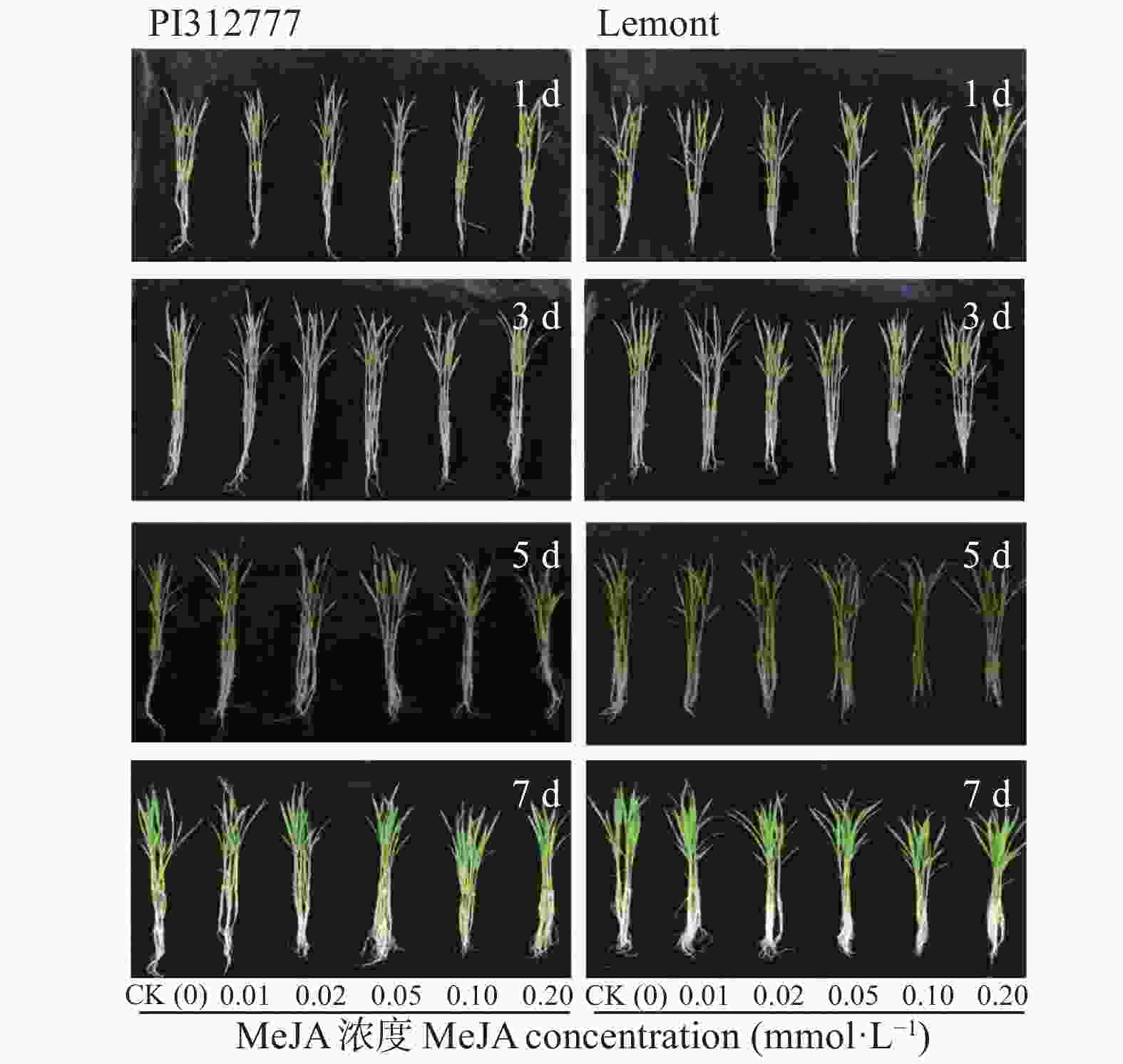
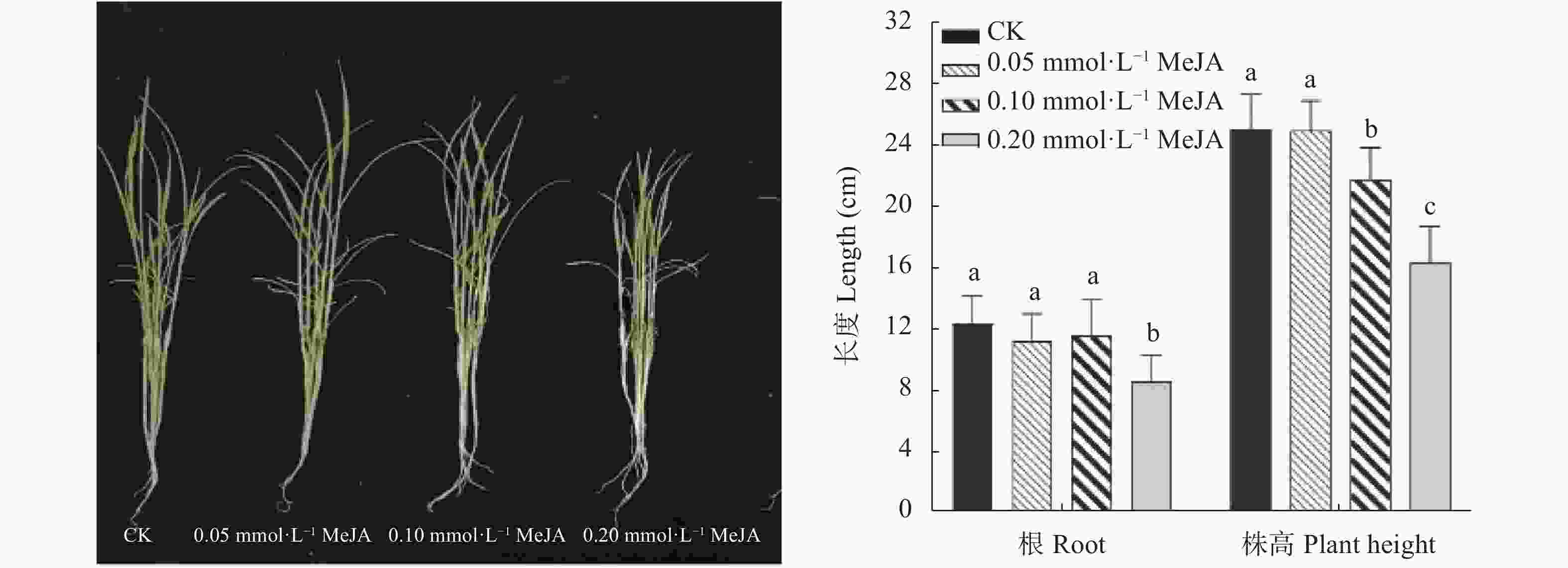
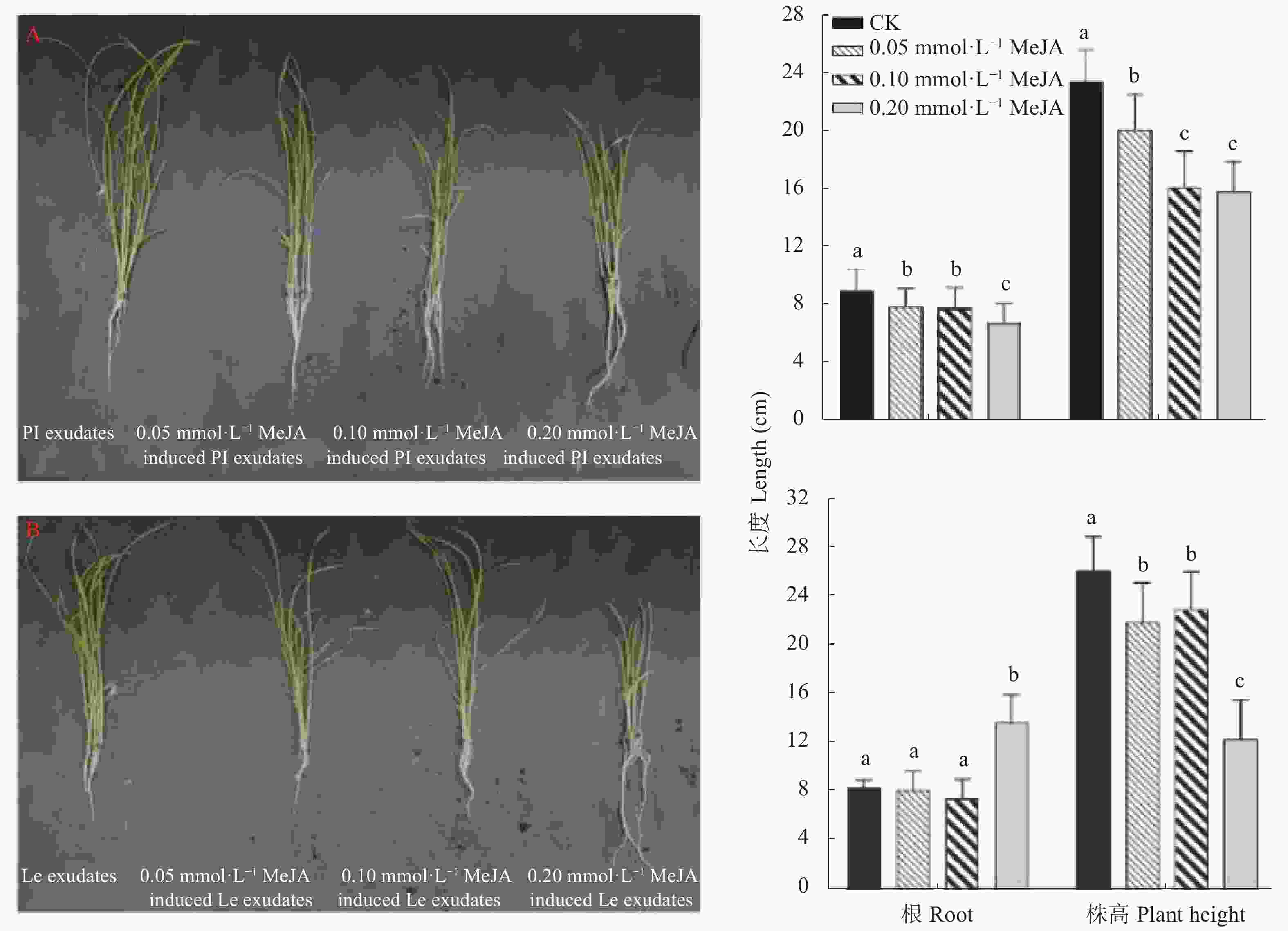
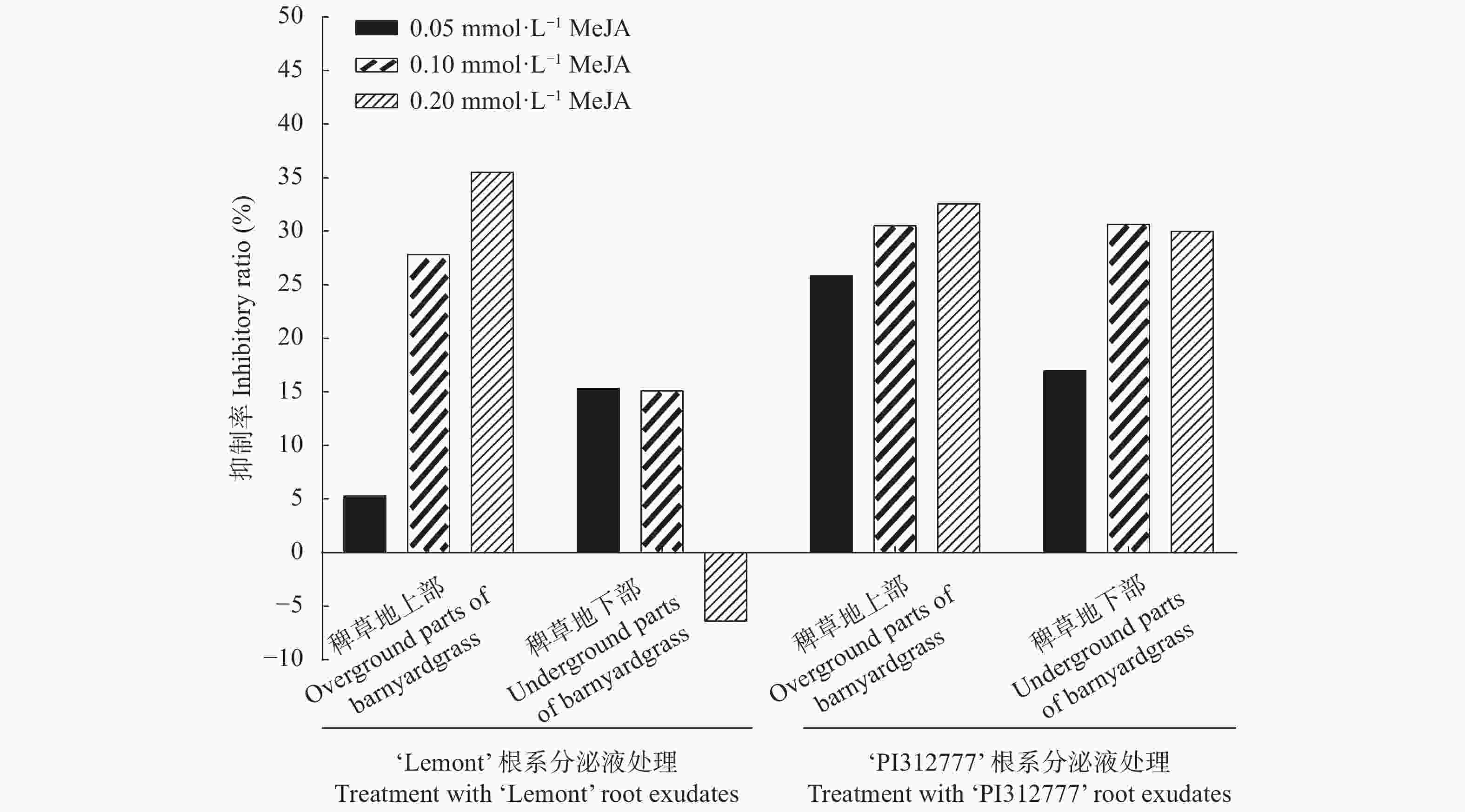
图 8 不同浓度茉莉酸甲酯处理下‘PI312777’ (PI)和‘Lemont’ (Le)水稻根系分泌液对稗草生长的影响
不同小写字母表示不同茉莉酸甲脂浓度间差异显著(P<0.05)。
Figure 8. Effect of root exudates of ‘PI312777’ (PI) and ‘Lemont’ (Le) rice induced by different concentrations of methyl jasmonate (MeJA) on barnyardgrass growth
Different lowercase letters indicate significant different among different MeJA concentrations at P<0.05.
表 1
本研究中采用的qPCR引物 Table 1. qPCR primers used in this study
基因名称
Gene name引物序列
Primer sequences (5′-3′)OsMYC2 F: GGGTTTTGTTATCTCCTCTCCT OsMYC2 R: AGAATAGGAGGAGAGGAAAGGT Os04g0397500 F: GAATCGCTGATAAGATGTAGCC Os04g0397500 R: CCCCATGCATTTTCCTTCTTTT Os02g0516800 F: GGAAGAGTCGCTGTTCTCC Os02g0516800 R: AGAAGAATAGTCGCGGATATCG Os03g0370250 F: AGCACCTCATAGTACATTGTCG Os03g0370250 R: CTCCTGCTAATCCTATCTGCAA Os03g0708800 F: CTCAAGAACAAGTTCCGCG Os03g0708800 R: GTAATCATGGGAATTGCTGTCC actin F: CTTCATAGGAATGGAAGCTGCGGGTA actin R: CGACCACCTTGATCTTCATGCTGCTA 表 2 ‘PI312777’和‘Lemont’水稻中OsMYB57基因启动子上的结合蛋白鉴定结果
Table 2. Identification of the proteins binding on the OsMYB57 gene promoter from rice varieties ‘PI312777’ and ‘Lemont’
水稻品种
Rice variety蛋白登录号
Protein_ID特异肽段数
Unique peptide number覆盖率
Coverage蛋白描述
Protein description基因
GenePI312777 LOC_Os10g42430.1 8 0.1558 basic helix-loop-helix protein 009 OsMYC2 LOC_Os04g32590.1 11 0.5410 transcription factor Os04g0397500 LOC_Os02g31160.1 11 0.4626 transcription factor Os02g0516800 LOC_Os03g25430.1 9 0.3926 transcription regulator Os03g0370250 LOC_Os03g50110.1 4 0.1733 transcription regulator Os03g0708800 Lemont LOC_Os04g32590.1 13 0.5137 transcription factor Os04g0397500 LOC_Os02g31160.1 13 0.5134 transcription factor Os02g0516800 LOC_Os03g25430.1 6 0.2122 transcription regulator Os03g0370250 -
[1] WESTON L A, DUKE S O. Weed and crop allelopathy[J]. Critical Reviews in Plant Sciences, 2003, 22(3/4): 367−389 [2] MALLIK A U. Allelopathy in forested ecosystems[M]// Allelopathy in Sustainable Agriculture and Forestry. New York: Springer New York, 2008: 363–386 [3] REIGOSA M S, GONZALEZY L, SOUTO X C, et al. Allelopathy in forest ecosystems[M]//Allelopathy in Ecological Agriculture and Forestry. Dordrecht: Springer Netherlands, 2000: 183–193 [4] GRANÉLI E, PAVIA H. Allelopathy in marine ecosystems[M]//Allelopathy. Dordrecht: Kluwer Academic Publishers, 2006: 415–431 [5] INDERJIT, DAKSHINI K M M. Algal allelopathy[J]. The Botanical Review, 1994, 60(2): 182−196 doi: 10.1007/BF02856576 [6] HE H B, WANG H B, FANG C X, et al. Barnyard grass stress up regulates the biosynthesis of phenolic compounds in allelopathic rice[J]. Journal of Plant Physiology, 2012, 169(17): 1747−1753 doi: 10.1016/j.jplph.2012.06.018 [7] LI J Y, ZHANG Q, YANG X Y, et al. A reappraisal of the content and the differences of phenolic acids between allelopathic and non-allelopathic rice accessions[J]. Allelopathy Journal, 2017, 40(1): 35−46 doi: 10.26651/2017-40-1-1064 [8] LI L L, ZHAO H H, KONG C H. (−)-Loliolide, the most ubiquitous lactone, is involved in barnyardgrass-induced rice allelopathy[J]. Journal of Experimental Botany, 2019, 71(4): 1540−1550 [9] KATO-NOGUCHI H, PETERS R J. The role of momilactones in rice allelopathy[J]. Journal of Chemical Ecology, 2013, 39(2): 175−185 doi: 10.1007/s10886-013-0236-9 [10] KONG C H, ZHAO H, XU X H, et al. Activity and allelopathy of soil of Flavone O-glycosides from rice[J]. Journal of Agricultural and Food Chemistry, 2007, 55(15): 6007−6012 doi: 10.1021/jf0703912 [11] 王海斌, 俞振明, 何海斌, 等. 不同化感潜力水稻化感效应与产量的关系[J]. 中国生态农业学报, 2012, 20(1): 75−79 doi: 10.3724/SP.J.1011.2012.00075WANG H B, YU Z M, HE H B, et al. Relationship between allelopathic potential and grain yield of different allelopathic rice accessions[J]. Chinese Journal of Eco-Agriculture, 2012, 20(1): 75−79 doi: 10.3724/SP.J.1011.2012.00075 [12] BATISH D R, SINGH H P, KAUR S. Crop allelopathy and its role in ecological agriculture[J]. Journal of Crop Production, 2001, 4(2): 121−161 doi: 10.1300/J144v04n02_03 [13] NARWAL S S, HAOUALA R. Role of allelopathy in weed management for sustainable agriculture[M]//Allelopathy. Berlin, Heidelberg: Springer Berlin Heidelberg, 2012: 217–249 [14] FANG C X, LI Y Z, LI C X, et al. Identification and comparative analysis of microRNAs in barnyardgrass (Echinochloa crusgalli) in response to rice allelopathy[J]. Plant, Cell & Environment, 2015, 38(7): 1368−1381 [15] CAMM E L, TOWERS G H N. Phenylalanine ammonia-lyase[J]. Phytochemistry, 1973, 12(5): 961−973 doi: 10.1016/0031-9422(73)85001-0 [16] 李兰兰, 母丹, 严雪, 等. OsPAL2;3对水稻化感抑制稗草能力的调控作用[J]. 作物学报, 2021, 47(2): 197−209 doi: 10.3724/SP.J.1006.2021.02034LI L L, MU D, YAN X, et al. Effect of OsPAL2;3 in regulation of rice allopathic inhibition on barnyardgrass (Echinochloa crusgalli L.)[J]. Acta Agronomica Sinica, 2021, 47(2): 197−209 doi: 10.3724/SP.J.1006.2021.02034 [17] FAROOQ M A, ISLAM F, YANG C, et al. Methyl jasmonate alleviates arsenic-induced oxidative damage and modulates the ascorbate-glutathione cycle in oilseed rape roots[J]. Plant Growth Regulation, 2018, 84(1): 135−148 doi: 10.1007/s10725-017-0327-7 [18] BI H H, ZENG R S, SU L M, et al. Rice allelopathy induced by methyl jasmonate and methyl salicylate[J]. Journal of Chemical Ecology, 2007, 33(5): 1089−1103 doi: 10.1007/s10886-007-9286-1 [19] DUBOS C, STRACKE R, GROTEWOLD E, et al. MYB transcription factors in Arabidopsis[J]. Trends in Plant Science, 2010, 15(10): 573−581 doi: 10.1016/j.tplants.2010.06.005 [20] LIU J Y, OSBOURN A, MA P D. MYB transcription factors as regulators of phenylpropanoid metabolism in plants[J]. Molecular Plant, 2015, 8(5): 689−708 doi: 10.1016/j.molp.2015.03.012 [21] DIAS A P, GROTEWOLD E. Manipulating the accumulation of phenolics in maize cultured cells using transcription factors[J]. Biochemical Engineering Journal, 2003, 14(3): 207−216 doi: 10.1016/S1369-703X(02)00225-5 [22] LUO J, BUTELLI E, HILL L, et al. AtMYB12 regulates caffeoyl quinic acid and flavonol synthesis in tomato: expression in fruit results in very high levels of both types of polyphenol[J]. The Plant Journal, 2008, 56(2): 316−326 doi: 10.1111/j.1365-313X.2008.03597.x [23] ROMMENS C M, RICHAEL C M, YAN H, et al. Engineered native pathways for high kaempferol and caffeoylquinate production in potato[J]. Plant Biotechnology Journal, 2008, 6(9): 870−886 doi: 10.1111/j.1467-7652.2008.00362.x [24] PAYYAVULA R S, SINGH R K, NAVARRE D A. Transcription factors, sucrose, and sucrose metabolic genes interact to regulate potato phenylpropanoid metabolism[J]. Journal of Experimental Botany, 2013, 64(16): 5115−5131 doi: 10.1093/jxb/ert303 [25] QIU J, GAO F H, SHEN G A, et al. Metabolic engineering of the phenylpropanoid pathway enhances the antioxidant capacity of Saussurea involucrata[J]. PLoS One, 2013, 8(8): e70665 doi: 10.1371/journal.pone.0070665 [26] DELUC L, BARRIEU F, MARCHIVE C, et al. Characterization of a grapevine R2R3-MYB transcription factor that regulates the phenylpropanoid pathway[J]. Plant Physiology, 2006, 140(2): 499−511 doi: 10.1104/pp.105.067231 [27] STRACKE R, ISHIHARA H, HUEP G, et al. Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling[J]. The Plant Journal, 2007, 50(4): 660−677 doi: 10.1111/j.1365-313X.2007.03078.x [28] FANG C X, YANG L K, CHEN W S, et al. MYB57 transcriptionally regulates MAPK11 to interact with PAL2;3 and modulate rice allelopathy[J]. Journal of Experimental Botany, 2020, 71(6): 2127−2141 doi: 10.1093/jxb/erz540 [29] GEALY D R, WAILES E J, ESTORNINOS JR L E, et al. Rice cultivar differences in suppression of barnyardgrass (Echinochloa crusgalli) and economics of reduced propanil rates[J]. Weed Science, 2013, 51(4): 601−609 [30] YOSHIDA S. Routine procedure for growing rice plants in culture solution[R]//Laboratory Manual for Physiological Studies of Rice. Los Baños: The International Rice Research Institute, 1976: 61–66 [31] LIVAK K J, SCHMITTGEN T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method[J]. Methods, 2001, 25(4): 402−408 doi: 10.1006/meth.2001.1262 [32] 秦利征, 赵全志, 李俊周. 水稻根系蛋白质提取方法的改良[J]. 分子植物育种, 2013, 11(4): 588−594QIN L Z, ZHAO Q Z, LI J Z. The modified method for rice root protein extraction[J]. Molecular Plant Breeding, 2013, 11(4): 588−594 [33] FANG C X, XIONG J, QIU L, et al. Analysis of gene expressions associated with increased allelopathy in rice (Oryza sativa L.) induced by exogenous salicylic acid[J]. Plant Growth Regulation, 2008, 57(2): 163−172 [34] CHUNG I M, KIM K H, AHN J K, et al. Screening of allelochemicals on barnyardgrass (Echinochloa crusgalli) and identification of potentially allelopathic compounds from rice (Oryza sativa) variety hull extracts[J]. Crop Protection, 2002, 21(10): 913−920 doi: 10.1016/S0261-2194(02)00063-7 [35] VOM ENDT D, KIJNE J W, MEMELINK J. Transcription factors controlling plant secondary metabolism: what regulates the regulators?[J]. Phytochemistry, 2002, 61(2): 107−114 doi: 10.1016/S0031-9422(02)00185-1 [36] OKSMAN-CALDENTEY K M, INZÉ D. Plant cell factories in the post-genomic era: new ways to produce designer secondary metabolites[J]. Trends in Plant Science, 2004, 9(9): 433−440 doi: 10.1016/j.tplants.2004.07.006 [37] NIU Y J, FIGUEROA P, BROWSE J. Characterization of JAZ-interacting bHLH transcription factors that regulate jasmonate responses in Arabidopsis[J]. Journal of Experimental Botany, 2011, 62(6): 2143−2154 doi: 10.1093/jxb/erq408 [38] 蔡昆争, 董桃杏, 徐涛. 茉莉酸类物质(JAs)的生理特性及其在逆境胁迫中的抗性作用[J]. 生态环境, 2006, 15(2): 397−404CAI K Z, DONG T X, XU T. The physiological roles and resistance control in stress environment of jasmonates[J]. Ecology and Environment, 2006, 15(2): 397−404 [39] ORTIGOSA A, FONSECA S, FRANCO-ZORRILLA J M, et al. The JA-pathway MYC transcription factors regulate photomorphogenic responses by targeting HY5 gene expression[J]. The Plant Journal, 2020, 102(1): 138−152 doi: 10.1111/tpj.14618 [40] FANG C X, YU Y, CHEN W S, et al. Role of allene oxide cyclase in the regulation of rice phenolic acids synthesis and allelopathic inhibition on barnyardgrass[J]. Plant Growth Regulation, 2016, 79(3): 265−273 doi: 10.1007/s10725-015-0131-1 [41] BHAVANAM S, STOUT M. Seed treatment with jasmonic acid and methyl jasmonate induces resistance to insects but reduces plant growth and yield in rice, Oryza sativa[J]. Frontiers in Plant Science, 2021, 12: 691768 doi: 10.3389/fpls.2021.691768 [42] BERTINI L, PALAZZI L, PROIETTI S, et al. Proteomic analysis of MeJA-induced defense responses in rice against wounding[J]. International Journal of Molecular Sciences, 2019, 20(10): 2525 [43] YANG Y C, DU Z M, ZHANG X P, et al. Effects of spraying methyl jasmonate on yield and grain quality of japonica rice during heading and flowering stage[J]. Crops, 2021, 37(2): 71−76 -





 下载:
下载: